isolate umap from scanpy to scv Split.by-like function for the umap to compare gene expression between ...
If you are searching about Unexplained change in UMAP - scanpy - scverse you've visit to the right page. We have 10 Pictures about Unexplained change in UMAP - scanpy - scverse like GitHub - adhiraj141092/SCANPY_GUI: A gui for single cell RNA-Seq data, Unexplained change in UMAP - scanpy - scverse and also split.by-like function for the umap to compare gene expression between. Here you go:
Unexplained Change In UMAP - Scanpy - Scverse
 discourse.scverse.org
discourse.scverse.org
Unexplained change in UMAP - scanpy - scverse
Umap Failed To Cluster The Cells · Issue #2386 · Scverse/scanpy · GitHub
 github.com
github.com
umap failed to cluster the cells · Issue #2386 · scverse/scanpy · GitHub
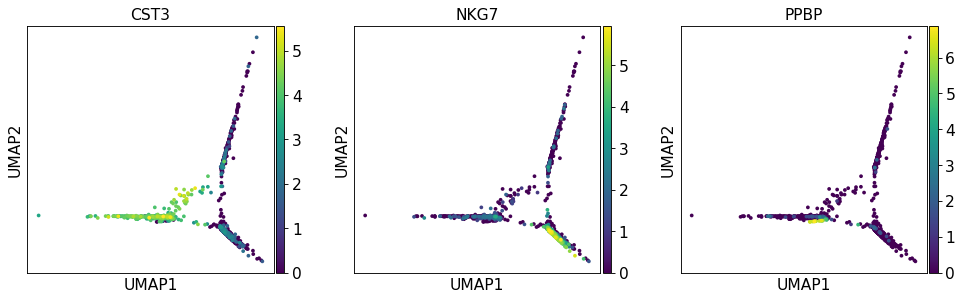
Wrong UMAP Output · Issue #2101 · Scverse/scanpy · GitHub
 github.com
github.com
Wrong UMAP output · Issue #2101 · scverse/scanpy · GitHub
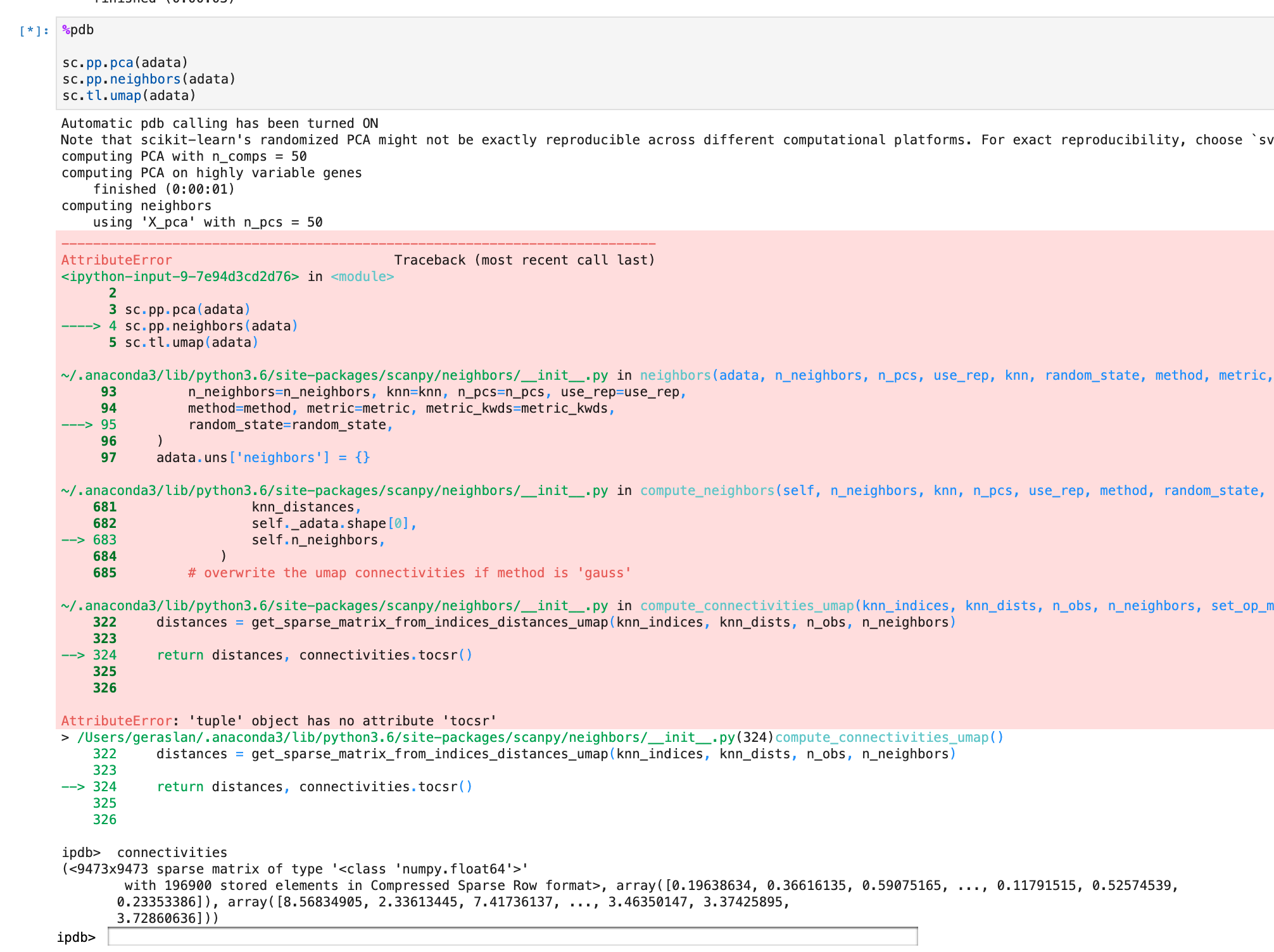
Preparations For UMAP 0.4 Release · Issue #779 · Scverse/scanpy · GitHub
 github.com
github.com
Preparations for UMAP 0.4 release · Issue #779 · scverse/scanpy · GitHub
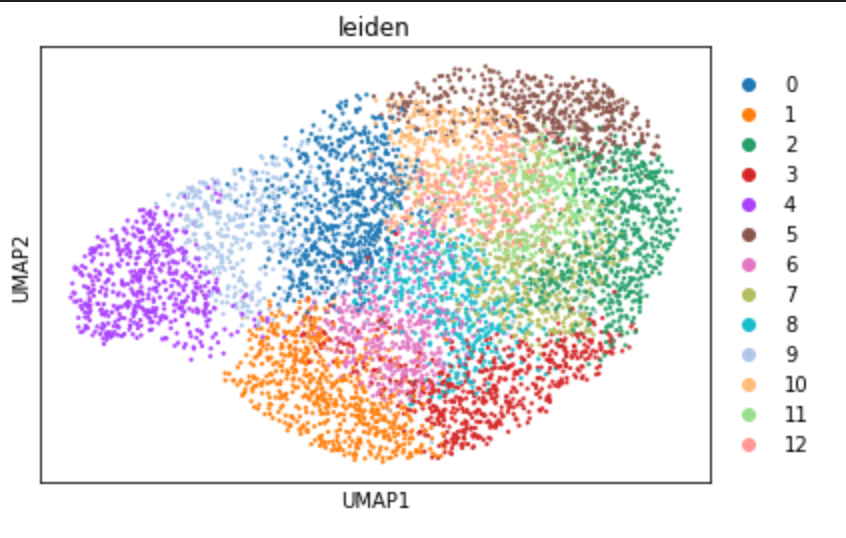
UMAP Goes Wrong On Scanpy V1.8.2 · Issue #2148 · Scverse/scanpy · GitHub
 github.com
github.com
UMAP goes wrong on scanpy v1.8.2 · Issue #2148 · scverse/scanpy · GitHub
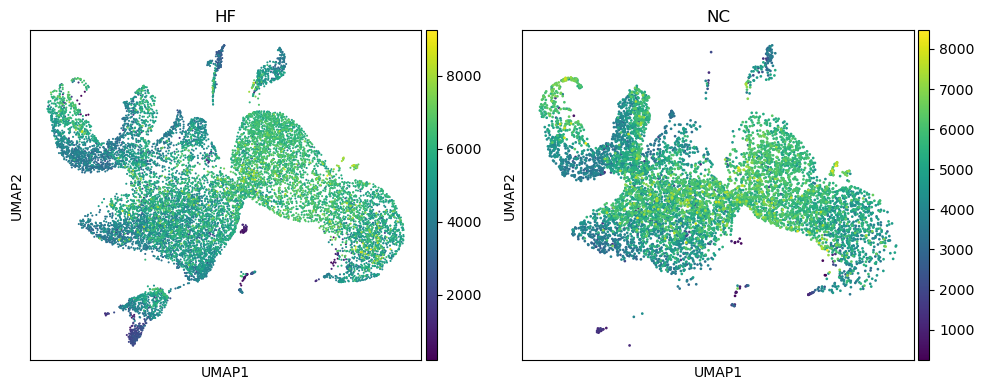
Split.by-like Function For The Umap To Compare Gene Expression Between
 github.com
github.com
split.by-like function for the umap to compare gene expression between ...
Problem In Spatial Umap Plot · Issue #2291 · Scverse/scanpy · GitHub
 github.com
github.com
Problem in Spatial Umap Plot · Issue #2291 · scverse/scanpy · GitHub
Sc.pl.umap(colors=['cell_type', 'other'], Legend_loc=['on Data', 'right
sc.pl.umap(colors=['cell_type', 'other'], legend_loc=['on data', 'right ...
GitHub - Adhiraj141092/SCANPY_GUI: A Gui For Single Cell RNA-Seq Data
 github.com
github.com
GitHub - adhiraj141092/SCANPY_GUI: A gui for single cell RNA-Seq data ...
Unexplained Change In UMAP - Scanpy - Scverse
 discourse.scverse.org
discourse.scverse.org
Unexplained change in UMAP - scanpy - scverse
Wrong umap output · issue #2101 · scverse/scanpy · github. Split.by-like function for the umap to compare gene expression between. Umap failed to cluster the cells · issue #2386 · scverse/scanpy · github